The Cyclic di-AMP Signaling Project
Updated Jan. 2023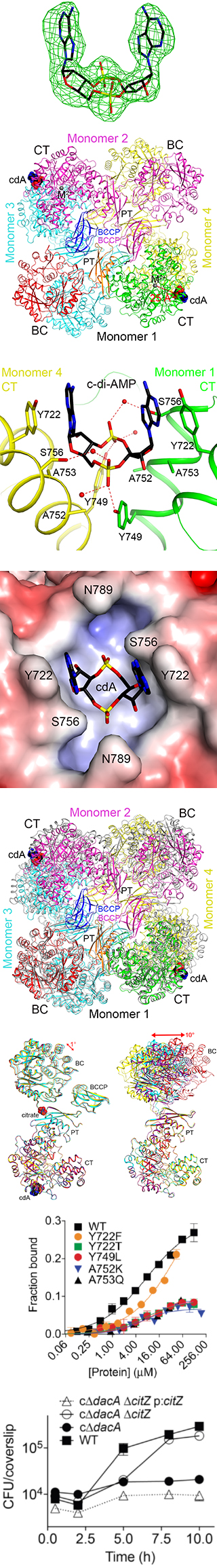
Cyclic di-AMP (c-di-AMP, cdA) was first discovered in 2008, and this second messenger has crucial roles in many bacterial processes, including central metabolism, cell wall metabolism, DNA repair, potassium homeostasis, osmotic regulation, sporulation, stress response, antibiotic resistance, biofilm formation and virulence. It is essential in many bacteria, and riboswitches and proteins have been identified as its sensors. It also has an important role in host-pathogen interactions.
Using a chemical proteomics approach, a set of proteins that interact with c-di-AMP was identified in Listeria monocytogenes. The biochemical and physiological effects of c-di-AMP binding to these target proteins were then characterized. The molecular basis for the binding was also determined.
Major findings from this project- The structure of Listeria monocytogenes pyruvate carboxylase (LmPC) in complex with c-di-AMP has been determined, the first structure of c-di-AMP bound to a protein target.
- c-di-AMP is bound at the dimer interface of the CT domain of LmPC, showing pi-stacking, hydrogen-bonding and van der Waals interactions with the enzyme.
- There are large conformational changes in LmPC upon c-di-AMP binding. The four monomers of LmPC assume a single conformation in the complex, while they have large variability in the free enzyme. This 'freezing' of the enzyme may be the molecular basis of the inhibitory effect of c-di-AMP.
- Mutations in the c-di-AMP binding site can abolish binding.
- Reduced levels of c-di-AMP in Listeria causes a metabolic imbalance, which affects the growth of this bacterium in host cells.
- PC enzyzmes from other bacteria that have a conserved binding site are also sensitive to c-di-AMP, such as that from Lactococcus lactis (LlPC).
Publications from this project
- K. Sureka,* P.H. Choi,* M. Precit, M. Delince, D.A. Pensinger, T.N. Huynh, A.R. Jurado, Y.A. Goo, M. Sadilek, A.T. Iavarone, J.-D. Sauer, L. Tong$ & J.J. Woodward.$ (2014). The cyclic dinucleotide c-di-AMP is an allosteric regulator of metabolic enzyme function. Cell, 158, 1389-1401. (*-equal first authors, $-co-corresponding authors)
- T.N. Huynh*, S. Luo*, D.A. Pensinger, J.-D. Sauer, L. Tong$ & J.J. Woodward.$. (2015). An HD-domain phosphodiesterase mediates cooperative hydrolysis of c-di-AMP to affect bacterial growth and virulence. Proc. Natl. Acad. Sci. USA, 112, E747-E756. (*-equal first authors, $-co-corresponding authors)
- P.H. Choi*, K. Surekha*, J.J. Woodward$ & L. Tong.$ (2015). Molecular basis for the recognition of cyclic-di-AMP by PstA, a PII-like signal transduction protein. MicrobiologyOpen, 4, 361-374. (*-equal first authors, $-co-corresponding authors)
- T.N. Huynh*, P.H. Choi*, K. Sureka, H.E. Ledvina, J. Campillo, L. Tong$ & J.J. Woodward.$ (2016). Cyclic di-AMP targets the cystathionine beta-synthase domain of the osmolyte transporter OpuC. Mol. Microbiol. 102, 233-243. (*-equal first authors, $-co-corresponding authors)
- A.P. McFarland, S. Luo, F. Ahmed-Qadri, M. Zuck, E.F. Thayer, Y.A. Goo, K. Hybiske, L. Tong & J.J. Woodward. (2017). Sensing of bacterial cyclic dinucleotides by the oxidoreductase RECON promotes NF-kB activation and shapes a proinflammatory antibacterial state. Immunity, 46, 433-445.
- A.T. Whiteley, N.E. Garelis, B.N. Peterson, P.H. Choi, L. Tong, J.J. Woodward & D.A. Portnoy. (2017). c-di-AMP modulates Listeria monocytogenes central metabolism to regulate growth, antibiotic resistance, and osmoregulation. Mol. Microbiol., 104, 212-233.
- P.H. Choi, T.M.N. Vu, H.T. Pham, J.J. Woodward, M.S. Turner & L. Tong. (2017). Structural and functional studies of pyruvate carboxylase regulation by cyclic-di-AMP in lactic acid bacteria. Proc. Natl. Acad. Sci. USA, 114, E7226-E7235.
- B.N. Peterson, M.K.M. Young, S. Luo, J. Wang, A.T. Whiteley, J.J. Woodward, L. Tong, J.D. Wang & D.A. Portnoy. (2020). (p)ppGpp and c-di-AMP homeostasis is controlled by CbpB in Listeria monocytogenes. mBio, 11, e01625-20.
- L. Tong$ & J.J. Woodward.$ (2020). Metabolic regulation by cyclic di-AMP signaling. in Microbial Cyclic Di-Nucleotide Signaling, Chapter 10, pp. 161-175. S.-H. Chou, N. Guiliani, V. Lee & U. Romling Eds. Springer, New York, Heidelberg, Dordrecht, London. ($-co-corresponding authors)
- A.J. Pollock, P.H. Choi, S.A. Zaver, L. Tong & J.J. Woodward. (2021). A rationally designed c-di-AMP Forster resonance energy transfer biosensor to monitor nucleotide dynamics. J. Bacteriol. 203, e0008021.
Funding for this project
- NIH R01AI116669 (PI: Woodward, 2015-2020)
© copyright 2019-2023, Liang Tong.