The Carnitine Acyltransferases Project
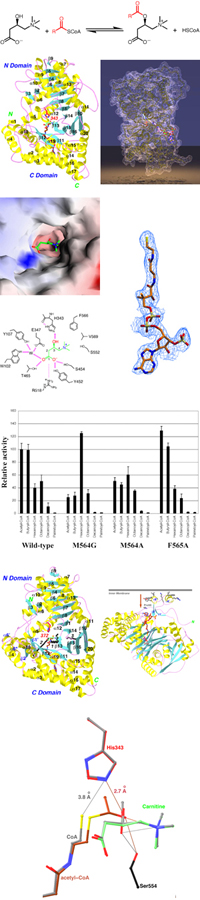
Carnitine acyltransferases catalyze the exchange between acyl-CoA and acylcarnitines. The enzymes can be divided into three sub-families based on their substrate preferences: carnitine acetyltransferases (CrAT), carnitine octanoyltransferases (CrOT), and carnitine palmitoyltransferases (CPT), with preferences for short-chain, medium-chain, and long-chain fatty acids, respectively.
The CPTs have crucial roles in the oxidation of long-chain fatty acids. L-CPT-I and M-CPT-I are located on the outer membrane of the mitochondria, and convert fatty acyl-CoAs to fatty acylcarnitines. Upon translocation into the mitochondria, CPT-II then converts them back to fatty acyl-CoAs, which can then undergo b-oxidation for energy production.
L-CPT-I is a target for the development of drugs against type 2 diabetes. At the same time, an agonist of this enzyme may be efficacious in the control of body weight and obesity
The sequences of the enzymes display signicant homology to each other (35% identity or better), but they do not show any recognizable homology to other proteins in the database.
Major findings from this project- The structure of mouse CrAT has been determined at 1.8 A resolution.
- The backbone fold of CRAT shows unexpected homology to chloramphenicol acetyltransferase (CAT). However, the quaternary structures of the two enzymes are different. CAT is a homo trimer, whereas CrAT contains two domains that mimics the two subunits of the CAT trimer.
- There is a tunnel that goes through the middle of the protein. Carnitine and CoA bind to opposite ends of this tunnel.
- The carboxyl group of carnitine is involved in an intricate network of hydrogen-bonding interactions.
- The CoA molecule is fully extended when bound to CrAT, in contrast to the folded conformation in the CAT complex.
- The positive charge on carnitine helps stabilize the oxyanion of the transition state - substrate-assisted catalysis.
- The Met564 residue of CrAT is a determinant of substrate preference. The M564G mutant has stronger activity with medium-chain substrates.
- The structure of mouse CrOT has been determined at 2.0A resolution.
- The structure of rat CPT-II has been determined at 1.9A resolution.
- An insert of two helices, with a hydrophobic surface patch, may mediate the attachment of CPT-II to the inner mitochondrial membrane.
- The P50H disease-causing mutation in CPT-II may affect its membrane localization.
- The structures of CrAT in ternary complex with its substrates (Michaelis complex) have been determined at up to 1.9A resolution.
Publications from this project
- G. Jogl & L. Tong. (2003). Crystal structure of carnitine acetyltransferase and implications for the catalytic mechanism and fatty acid transport. Cell 112, 113-122. Reprint(PDF)
- S. Gobin, L. Thuillier, G. Jogl, A. Faye, L. Tong, M. Chi, J.-P. Bonnefont, J. Girard, C. Prip-Buus. (2003). Functional and structural basis of carnitine palmitoyltransferase 1A deficiency. J. Biol. Chem. 278, 50428-50434. Reprint(PDF)
- Y.-S. Hsiao, G. Jogl & L. Tong. (2004). Structural and biochemical studies of the substrate selectivity of carnitine acetyltransferase. J. Biol. Chem. 279, 31584-31589. Reprint(PDF)
- G. Jogl, Y.-S. Hsiao & L. Tong. (2004). Structure and function of carnitine acyltransferases. Ann. N.Y. Acad. Sci. 1033, 17-29. Reprint(PDF)
- G. Jogl, Y.-S. Hsiao & L. Tong. (2005). Crystal structure of mouse carnitine octanoyltransferase and molecular determinants of substrate selectivity. J. Biol. Chem. 280, 738-744. Reprint(PDF)
- Y.-S. Hsiao, G. Jogl, V. Esser & L. Tong. (2006). Crystal structure of rat carnitine palmitoyltransferase II (CPT-II). Biochem. Biophys. Res. Commun. 346, 974-980. Reprint(PDF)
- Y.-S. Hsiao, G. Jogl & L. Tong. (2006). Crystal structures of murine carnitine acetyltransferase in ternary complexes with its substrates. J. Biol. Chem. 281, 28480-28487. Reprint(PDF)
Funding for this project
- NIH R01 DK67238
© copyright 2003-2017, Liang Tong.