The 3-Methylcrotonyl-CoA Carboxylase (MCC) Project
3-Methylcrotonyl-CoA carboxylase (MCC) catalyzes the biotin-dependent carboxylation of 3-methylcrotonyl-CoA. MCC is essential for the catabolism of the amino acid leucine. In Pseudomonas organisms, MCC is also involved in the metabolism of terpenoids.
Deficiencies of MCC activity in humans are linked to the disease methylcrotonylglycinuria (MCG) and other diseases. MCC deficiency is one of the most common metabolic disorders.
MCC holoenzyme is a 750-kD a6b6 dodecamer. The a subunit is known to contain the biotin carboxylase (BC) and biotin carboxyl carrier protein (BCCP) domains, and the b subunit supplies the carboxyltransferase (CT) activity.
MCC has strong sequence conservation with PCC (42 and 34% sequence identity for the a and b subunits, respectively). Both are 750-kD a6b6 dedecamers, and both are metabolic enzymes functioning in the mitochondrial matrix.
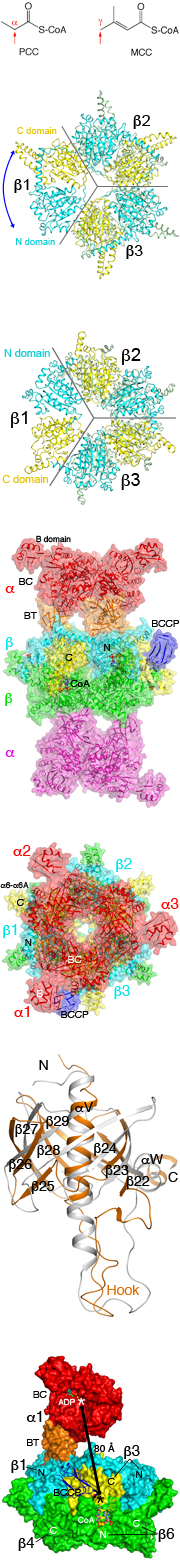
One difference between MCC and PCC is the site of carboxylation in the substrate. PCC carboxylates the a carbon of the organic acid (esterified to CoA), while MCC is active towards the g carbon of an a-b unsaturated acid.
Major findings from this project- The crystal structure of the b6 hexameric core of Pseudomonas MCC has been determined at 1.5A resolution.
- The positions of the N and C domains of the b subunit in MCC are swapped relative to those of the PCC b subunit.
- The N and C domains of the b subunit have the same backbone fold (crotonase fold, or b-b-a superhelix). The swapping of the domain positions is equivalent to a 60 degree pseudo rotation of the b6 hexamer.
- The crystal structures of the 750-kD a6b6 holoenzyme of Pseudomonas MCC have been determined at 2.9 and 3.5A resolution for the free enzyme and the CoA complex, respectively.
- An EM reconstruction of this enzyme, at 13 A resolution, shows the same structure.
- Despite their strong sequence conservation, the overall architecture of MCC is strikingly different from that of PCC.
- The holoenzyme has 32 symmetry, with a central b6 hexameric core.
- The a subunits are arranged as trimers in the holoenzyme, on both ends of the b6 core. This is in sharp contrast to the PCC holoenzyme, where the a subunits are arranged as monomers on both ends of the core.
- The overall structure of the BT domain is similar to that in PCC. However, the hook of the BT domain has a different conformation, and consequently the relative positioning of the a subunit relative to the b subunit is different in MCC compared to PCC.
- The active sites of BC and CT are separated by 85A in the holoenzyme, indicating that the entire BCCP domain must translocate during catalysis.
- BCCP and its biotin are located in the CT active site in the structure, providing insight into their involvement in the carboxyltransfer reaction.
- The structure of the holoenzyme provides a molecular basis for understanding the large collection of disease causing mutations in MCC.
- The structure of the 750kD holoenzyme of geranyl-CoA carboxylase (GCC) has been determined.
- The overall structure of the holoenzyme is remarkably similar to that of MCC.
- The presence of a Gly residue in the active site allows GCC to accommodate the larger geranyl group of the substrate. This residue is replaced with a Phe in MCC.
- The structures of MCC, GCC and PCC provide evidence in support of two distinct lineages of biotin-dependent acyl-CoA carboxylases, one carboxylating the alpha carbon of a saturated organic acid and the other carboxylating the gamma carbon of an alpha-beta unsaturated acid.
Publications from this project
- C.S. Huang, P. Ge, Z.H. Zhou & L. Tong. (2012). An unanticipated architecture of the 750kDa a6b6 holoenzyme of 3-methylcrotonyl-CoA carboxylase. Nature, 481, 219-223. Reprint(PDF)
- A.R. Jurado,* C.S. Huang,* X. Zhang, Z.H. Zhou & L. Tong. (2015). Structure of the 750 kD a6b6 holoenzyme of geranyl-CoA carboxylase and molecular basis for its substrate selectivity. Nature Commun. 6, 8986. (*-equal first authors) Reprint(PDF)
Funding for this project
- NIH R01 DK67238
© copyright 2012-2017, Liang Tong.