Enzymes involved in NAD+ metabolism
NAD+ is well-known as a cofactor for oxidation-reduction reactions. NAD+ is also a substrate for several important biochemical reactions - histone/protein deacetylation by the Sir2 family of enzymes, ADP-ribosylation of proteins (for example those catalyzed by the PARPs), and formation of cyclic ADP-ribose (important for calcium signaling). A common feature of these three reactions is that the glycosidic bond between nicotinamide and ribose is broken, destroying the NAD+ molecule. Cells need to replenish these NAD+ molecules through biosynthesis or other means.
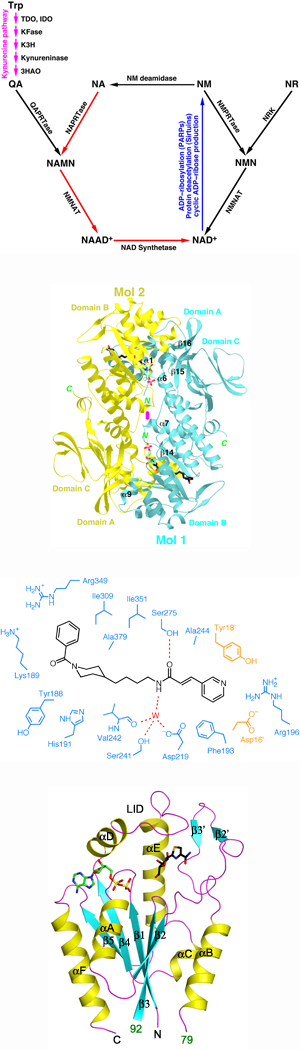
The biosynthesis of NAD+ can occur through several different pathways. The de novo pathway in eukaryotes uses Trp as the precursor. The kynurenine pathway converts Trp to quinolinic acid (QA), and QAPRTase converts QA to the mononucleotide NAMN. NAMN is then converted to the dinucleotide and amidated to produce NAD+. The salvage pathways can use either nicotinic acid (NA) or nicotinamide (NM) as the substrate. With NA, an NAPRTase converts it to NAMN. With NM, an NMPRTase converts to the mononucleotide NMN. NM is the breakdown product of NAD+ in the three reactions discussed above. Recently, another pathway of NAD+ biosynthesis was identified, using nicotinamide riboside (NR) as the precursor. NR kinase (NRK) converts NR to NMN. NRK is also involved in the activation of anticancer agents such as tiazofurin.
Cancer cells have elevated ADP-ribosylation activity, and it has been shown that blocking NAD+ biosynthesis can lead to apoptosis of these cells while having little effect against normal cells. A small molecule, FK866 (now known as APO866), is a potent inhibitor of NMPRTase and is currently in phase II clinical trials against some forms of cancers. NMPRTase has a crucial role in the salvage pathway using NM, the breakdown product of NAD+, as the precursor.
In yeast, NM is hydrolyzed to NA by Pnc1 (NM deamidase), which is linked to longevity in this organism.
Major findings from this project- The structures of human and mouse NMPRTase, free enzyme and complex with NMN or FK866, have been determined at up to 2.1 A resolution.
- NMPRTase is a tightly associated dimer, and the active site is located at the dimer interface.
- Asp219 is a determinant of the substrate selectivity between NM and NA by NMPRTase.
- FK866 is bound in a tunnel at the dimer interface, near the active site.
- FK866 is a competitive inhibitor of the NM substrate. Its apparent non-competitive inhibition in kinetic experiments is probably due to its tight-binding behavior.
- FK866 is specific for NMPRTase as the binding tunnel exists only in this enzyme.
- The crystal structures of human NRK, in complex with NMN or tiazofurin, have been determined at up to 1.5 A resolution.
- The crystal structures of Xanthomonas campestris TDO, the first enzyme of the kynurenine pathway, has been determined at up to 1.6 A resolution, alone and in complex with the substrate tryptophan.
- J.A. Khan, X. Tao, L. Tong. (2006). Molecular basis for the inhibition of human NMPRTase, a novel target for anticancer agents. Nature Struct. Mol. Biol. 13, 582-588. Reprint(PDF)
- F. Forouhar, J.L.R. Anderson, C.G. Mowat, S.M. Vorobiev, A. Hussain, M. Abashidze, C. Bruckmann, S.J. Thackray, J. Seetharaman, T. Tucker, R. Xiao, L.-C. Ma, L. Zhao, T.B. Acton, G.T. Montelione, Chapman, S.K., L. Tong. (2007). Molecular insights into substrate recognition and catalysis by tryptophan 2,3-dioxygenase. Proc. Natl. Acad. Sci. USA, 104, 473-478. Reprint(PDF)
- J.A. Khan, F. Forouhar, X. Tao, L. Tong. (2007). Nicotinamide adenine dinucleotide metabolism as an attractive target for drug discovery. Expert Opin. Ther. Targets, 11, 695-705. Reprint(PDF)
- J.A. Khan, S. Xiang, L. Tong (2007). Crystal structure of human nicotinamide riboside kinase. Structure, 15, 1005-1013. Reprint(PDF)
- NIH
© copyright 2007-2017, Liang Tong.