
- Home
- Research
Enzymes Involved in Fatty Acid/Carbohydrate Metabolism Acetyl-CoA Carboxylase (ACC) Propionyl-CoA Carboxylase (PCC) Methylcrotonyl-CoA Carboxylase (MCC) Pyruvate Carboxylase (PC) Carnitine acyltransferases AMP-activated Protein Kinase (AMPK/SNF1) ATP-citrate Lyase (ACLY) Malic Enzyme (ME)
Proteins Involved in mRNA Processing and/or Pol II Transcription Canonical pre-mRNA 3'-end Processing Histone pre-mRNA 3'-end Processing Integrator A Novel Quality Control Mechanism for 5'-end Capping 5'-3' Exoribonucleases (XRNs)
Additional Projects Cyclic di-AMP Signaling Separase Enzymes Involved in NAD Metabolism Crystallography Method/Software Development
Previous Projects Toll/Interleukin-1 Receptor (TIR) Domain BEACH Domain HCMV Protease
p38 MAP Kinase SH2 Domain HIV-2 Protease Human Renin Sindbis Virus Capsid Protein Human ras p21 - Publications
- Structures
- Software
- Members
- Pictures
- Links
- Contact
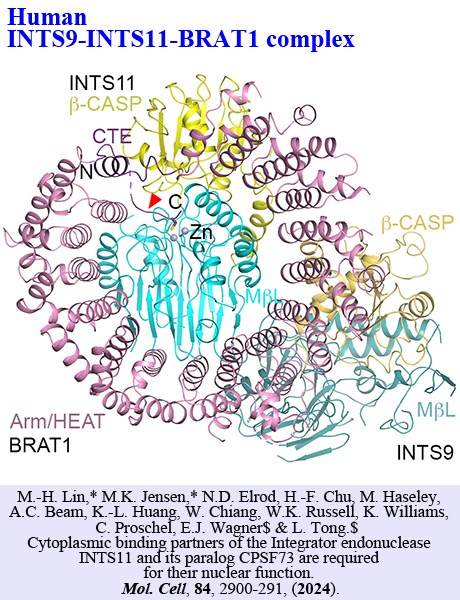 We use structural biology techniques (both cryo-EM and X-ray crystallography)
to elucidate the mechanism
and function of biological macromolecules.
We use structural biology techniques (both cryo-EM and X-ray crystallography)
to elucidate the mechanism
and function of biological macromolecules.
