
|
|

|
|
Brief Description of our ResearchWe use structural biology techniques (both cryo-EM and X-ray crystallography) to elucidate the mechanism and function of biological macromolecules.A major focus of our research is on proteins involved in pre-mRNA and snRNA 3'-end processing. Most eukaryotic mRNA precursors must undergo cleavage and polyadenylation in their 3'-ends before they can function as mRNAs. This canonical processing machinery contains more than 16 protein factors, which form several complexes (CPSF, CstF). CPSF contains two sub-complexes, mPSF (mammalian polyadenylation specificity factor) and mCF (mammalian cleavage factor). Replication-dependent histone pre-mRNAs are cleaved at their 3'-ends but not polyadenylated, and a distinct machinery is required for this processing. This machinery is the U7 snRNP, and histone pre-mRNA cleavage complex (HCC) is equivalent to the mCF in the canonical machinery. The goal of our research is to understand the molecular basis of this important event. We will produce structures of the protein subunits, protein-protein and protein-RNA sub-complexes, and the entire machinery, and carry out functional studies to assess the structural information. Our recent studies unexpectedly led to the identification of an mRNA 5'-end capping quality control mechanism. We have identified the enzymes (Rai1, Dxo1, DXO) that play a central role in this mechanism. We are characterizing their biochemical properties and deciphering their physiological functions in yeast as well as mammalian cells. Another area of our research is on enzymes that are involved in fatty acid and/or carbohydrate metabolism. These include acetyl-coenzyme A carboxylase (ACC), carnitine acyltransferase, AMP-activated protein kinase (AMPK), ATP-citrate lyase (ACLY) and others. These enzymes are important targets for drug discovery against obesity, diabetes and other human diseases. The goals of our research are to produce structural information on these enzymes and to understand their functions at the molecular level. The structural information will also lay the foundation for drug discovery against these targets. |
Structure Gallery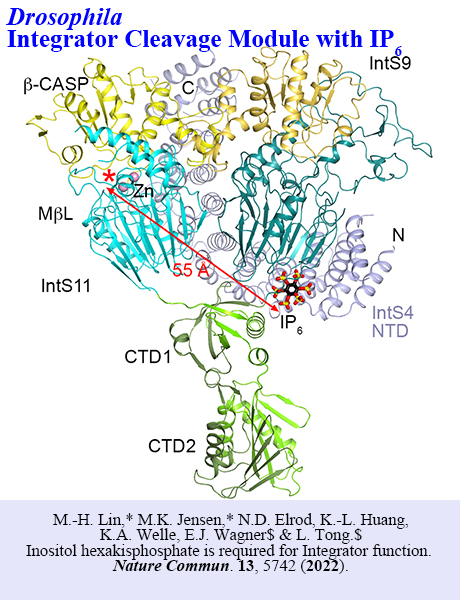
Recent PublicationsInositol hexakisphosphate is required for Integrator function. Nature Commun. 13, 5742 (2022). Y. Sun,* Y. Zhang,* W.S. Aik, X.-C. Yang, W.F. Marzluff, T. Walz$, Z. Dominski$ & L. Tong.$ Structure of an active human histone pre-mRNA 3'-end processing machinery. Science, 367, 700-703 (2020). J. Wei, S. Leit, J. Kuai, E. Therrien, S. Rafi, H.J. Harwood Jr, B. DeLaBarre & L. Tong. An allosteric mechanism for potent inhibition of human ATP-citrate lyase. Nature, 568, 566-570 (2019). |

|