
The Malic Enzyme (ME) Project
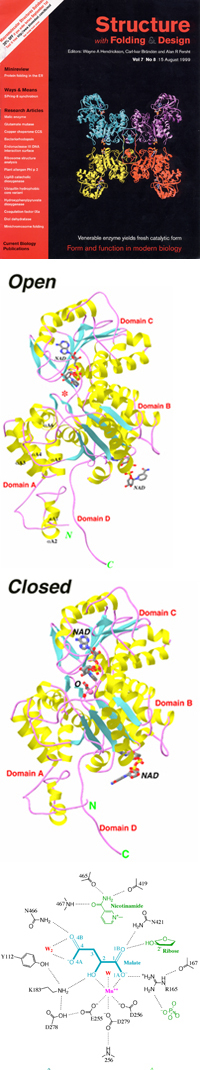
Malic enzymes catalyze the oxidative decarboxylation of malate to pyruvate
and CO2, with the concomitant reduction of the cofactor NAD(P)+ to NAD(P)H.
A divalent cation is required for this catalysis.
Malic enzymes have been found in most living organisms, from E. coli to man. The
amino acid sequences of malic enzymes are highly conserved, but they share little
sequence homology to other proteins.
Major findings from this project
- Malic enzymes belong to a new class of oxidative decarboxylases
- The divalent cation is bound by the side chains of three conserved
acid residues from the protein - Glu255, Asp256, Asp279. The substrate
malate provides two ligands, and the sixth ligand is a solvent water.
- Lanthanide ions are potent inhibitors of the enzyme and compete
for the cation binding site.
- An open and closed conformation of the enzyme have been observed.
- Conserved residues Tyr112 and Lys183 are important for the catalysis
by the enzyme.
- A second binding site for NAD+ may be the binding site for ATP,
an allosteric inhibitor of the enzyme.
Publications from this project
- G. Bhargava, S. Mui, S. Pav, H. Wu, G. Loeber & L. Tong. (1999). Preliminary crystallographic
studies of human mitochondrial NAD(P)+-dependent malic enzyme. J. Struct. Biol. 127, 72-75.
Reprint(PDF)
- Y. Xu, G. Bhargava, S. Mui, S. Pav, G. Loeber, H. Wu & L. Tong. (1999). Crystal structure of
human mitochondrial NAD(P)+-dependent malic enzyme: a new class of oxidative decarboxylases.
Structure, 7, 877-889.
Reprint(PDF)
- Z. Yang, D.L. Floyd, G. Loeber & L. Tong. (2000). Structure of closed form of malic enzyme and
implications for catalytic mechanism. Nature Struct. Biol. 7, 251-257.
Reprint(PDF)
- W.-Y. Chou, H.-P. Chang, C.-H. Huang, C.-C. Kuo, L. Tong & G.-G. Chang. (2000).
Characterization of the functional role of Asp141, Asp194, and Asp464 residues in the Mn2+-L-
malate binding of pigeon liver malic enzyme. Protein Sci. 9, 242-251.
Medline
- Z. Yang, R. Batra, D.L. Floyd, H.-C. Hung, G.-G. Chang & L. Tong. (2000). Potent and
competitive inhibition of malic enzymes by lanthanide ions. Biochem. Biophys. Res. Commun. 274,
440-444.
Medline
-
H.-C. Hung, G.-G. Chang, Z. Yang & L. Tong. (2000). Slow binding of metal ions to pigeon liver malic enzyme: a general case. Biochem. 39, 14095-14102.
-
Z. Yang & L. Tong. (2000). Structural studies of a human malic enzyme. Protein and Peptide Lett. 7, 287-296.
-
M.A. Rishavy, Z. Yang, L. Tong & W.W. Cleland. (2001).
Determination of the mechanism of human malic enzyme with natural
and alternate dinucleotides by isotope effects. Arch. Biochem. Biophys.
396, 43-48.
-
Z. Yang, H. Zhang, H.-C. Hung, C.-C. Kuo, L.-C. Tsai, H.S. Yuan,
W.-Y. Chou, G.-G. Chang & L. Tong. (2002). Structural studies
of pigeon cytosolic NADP+-dependent malic enzyme.
Protein Science, 11, 332-341.
Reprint(PDF)
-
Z. Yang, C.W. Lanks & L. Tong. (2002). Molecular mechanism for the regulation of human
mitochondrial NAD+-dependent malic enzyme by fumarate and ATP. Structure,
10, 951-960.
Reprint(PDF)
-
X. Tao, Z. Yang & L. Tong. (2003).
Crystal structures of substrate complexes of
malic enzyme and insights into the catalytic mechanism.
Structure, 11, 1141-1150.
Reprint(PDF)
-
G.-G. Chang & L. Tong. (2003).
Structure and function of malic enzymes, a new class
of oxidative decarboxylases.
Biochem. 42, 12721-12733.
Reprint(PDF)
-
C.-W. Kuo, H.-C. Hung, L. Tong & G.-G. Chang. (2004).
Metal-induced reversible structural interconversion of
human mitochondrial NAD(P)+-dependent malic enzyme.
Proteins, 54, 404-411.
-
W.-C. Hsu, H.-C. Hung, L. Tong & G.-G. Chang. (2004).
Dual functional roles of ATP in the human
mitochondrial malic enzyme.
Biochem. 43, 7382-7390.
Funding for this project
-
NSF MCB 99-74700 (1999-2003)
© copyright 2000-2026, Liang Tong.


